Marko Vendelin, Martin Laasmaa, Mari Kalda, Jelena Branovets, Niina Karro, Karina Barsunova, Rikke Birkedal
PLoS Comput Biol. 2020 Dec 22;16(12):e1008475.
- PMID: 33351800
- PMCID: PMC7787677
- DOI: 10.1371/journal.pcbi.1008475
Abstract
Biological measurements frequently involve measuring parameters as a function of time, space, or frequency. Later, during the analysis phase of the study, the researcher splits the recorded data trace into smaller sections, analyzes each section separately by finding a mean or fitting against a specified function, and uses the analysis results in the study. Here, we present the software that allows to analyze these data traces in a manner that ensures repeatability of the analysis and simplifies the application of FAIR (findability, accessibility, interoperability, and reusability) principles in such studies. At the same time, it simplifies the routine data analysis pipeline and gives access to a fast overview of the analysis results. For that, the software supports reading the raw data, processing the data as specified in the protocol, and storing all intermediate results in the laboratory database. The software can be extended by study- or hardware-specific modules to provide the required data import and analysis facilities. To simplify the development of the data entry web interfaces, that can be used to enter data describing the experiments, we released a web framework with an example implementation of such a site. The software is covered by open-source license and is available through several online channels.
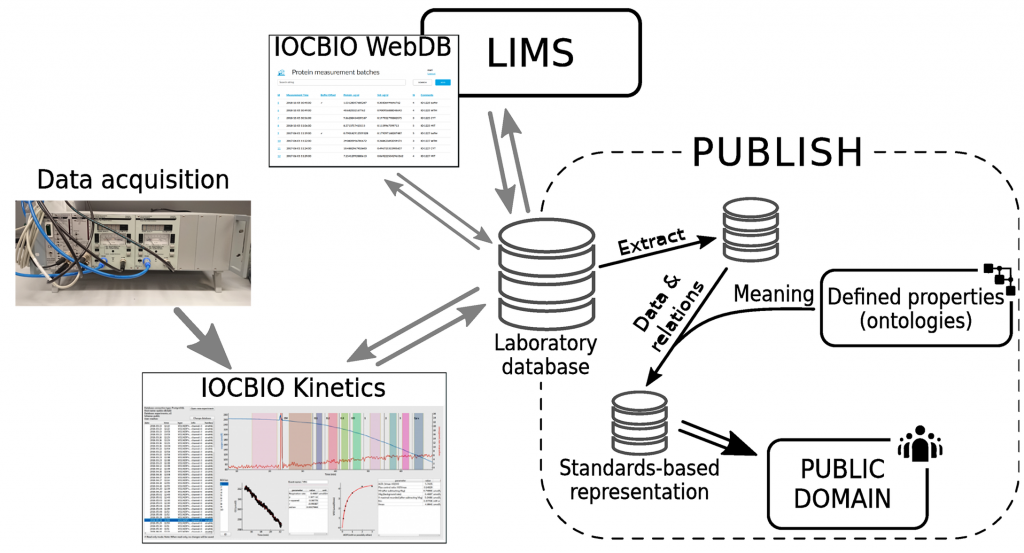
More about software: IOCBIO Kinetics and IOCBIO WebDB.